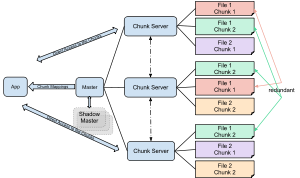
We do however update the totIonCurrent header value upon export with the sum of intensities, so the tic of the re-read mzXML file is identical to the tic calculated as the sum of intensities per spectrum. To test I exported the CDF above as mzXML file and compared the tic of that with the tic of the original.  
Regarding the differences you see between CDF and mzXML, could it be explained by the same fact? I do not have data in both CDF and mzXML format to check, unfortunately. It might well be that this is specific for this file (no idea how this was generated or manipulated) - I haven't checked any other CDF files since I have none at hand.ĭoes this explain the differences that you see too? Plot(intensity( tic_chr), tic_calc, pch = 16, col = "#00000080 ")Īs you see, the reported total ion current is completely different to the sum of intensities per spectrum. # Finally extract a TIC tic_chr <- chromatogram( od, aggregationFun = "sum ") 
# Calculate the TIC tic_calc <- unlist(spectrapply( od, function( z) sum( z intensity))) Od <- readMSData(system.file( "cdf/ko15.CDF ", package = "msdata "), mode = "onDisk ")
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
